Nov. 1, 2013
3 Bullets:
- There exist copious amounts of public research data that can reveal new biological information if they are integrated and analyzed.
- One of ISB’s specialties is the ability to apply systems approaches to develop methods to integrate and analyze data.
- In the latest publication, ISB demonstrates an open-access computational strategy that can help any researcher capitalize on large data sets.
By Dr. Nitin Baliga
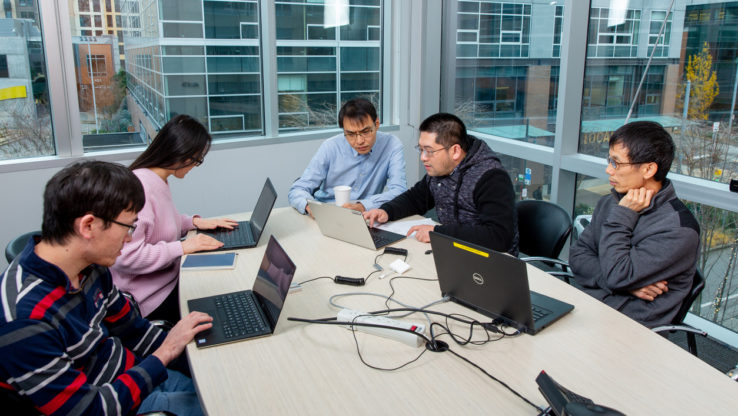
Systems biology promises to open a new world in human healthcare by integrating high throughput (big data) experiments to quickly discover novel biology and thus develop more effective treatment options. However, systems biology studies are often challenged by the problem of scale. In other words, how does one get from an abstract model encompassing all genes to specific experiments that reveal novel insight into biological mechanisms underlying health and disease. A team led by John Aitchison and Nitin Baliga has developed a template to discover novel biology from large amount of data that are now available for numerous organisms of medical and environmental importance. The work was published today in the journal Nucleic Acids Research.
Sam Danziger, a bioinformatics scientist working jointly under Drs. Aitchison and Baliga at ISB, developed a computational strategy to reconstruct the network of gene interactions within yeast using publicly available data from about 1500 experiments that were performed by different researchers across the world. Danziger and co-workers used this model to investigate how yeast cells regulate peroxisome function. Peroxisomes are medically important organelles that have been implicated in metabolic disorders and also the human innate immune response. Although a great deal of peroxisome biology has been characterized, much of the early events and mechanisms responsible for induction of this organelle are poorly understood. By iteratively refining the model, Danziger and co-workers discovered new genes and interactions across the varied hierarchies of genetic information processing to regulate peroxisome function, providing new leads for further studies in more complex mammalian cells.
Most notably, this study provides a template for molecular systems approaches to exploit large public data sets, which are increasingly available for varied biological systems. Applications include fundamental research into genetic regulation, identification of drug targets in diseased or pathogen-infected cells and engineering microorganisms for remediation or production of biomaterials.



 isbscience.org/research/how-isbs-systems-approach-finds-new-biological-insight-from-existing-big-data-sets/
isbscience.org/research/how-isbs-systems-approach-finds-new-biological-insight-from-existing-big-data-sets/